Bhattacharya lab hosts 4-H Rutgerscience Saturday on Coral Reef Genomics

About 500 million people worldwide depend on coral reefs for food, building materials, tourist income or coastal reef protection. © XL-Catlin-Seaview-Survey
Science agent and 4–H event coordinator Janice McDonnell welcomed 25 students between the ages of 10 – 13 years, who arrived from nearby locations and as far away as Bergen, Somerset, Mercer, Middlesex, Warren, and Hunterdon counties.

Students fill their test containers with water of various acidities.
During a plenary presentation Debashish Bhattacharya explained why Coral Reefs are an important ecosystem and their economic significance to mankind. The students learned about the major threats like coral bleaching, which occurs when warm water leads to expulsion of the algae that live inside the calcium carbonate skeleton, and can lead to coral death. The students learned that analysis of the coral genome uses information from the coral's entire DNA to figure out how they live, how they deal with stress, and today as the focus, how polyps build their stone houses (corallites) that make up reefs; that is, do biomineralization. Dr. Bhattacharya also talked about the importance of scientists making collaborative efforts to get answers to complex questions.

4–H students observe their experiments.
Graduate student Alexander Shumaker led an experiment where students tested the effect of water acidity on calcium carbonate (in the form of chalk). Coral skeletons are made of another form of calcium carbonate referred to as aragonite. While waiting for the results of their chemical reactions, the students learned basic terminology and concepts in bioinformatics.

Students taking a tour of the lab.
Starting with DNA sequencers and databases to software tools that help to analyze and compare patterns of gene occurrence in genomes. Alex presented photos from his doctoral expedition where he collected samples from coral reef sites, and how the samples need to be preserved so they could eventually be used for genome analysis in the lab. Concluding this session the students went on a lab tour where they became familiar with each necessary step along the way from holding a sample to preparing scientific data for publishing (e.g., view Montipora capitata samples in a -80 refrigerator, wet bench preparations, the sequencing machine).
In a parallel session, research associate Huan Qiu and graduate student Fatima Foflonker led the students to a computer room.

Participants analyze coral DNA in the
NCBI database.
By looking at the Foflonker family tree, the students learned that the further two people are apart from each other in the tree (of life), the less closely they are related. At the DNA level, the differences in DNA sequences measures this difference, for example, between humans and other species that can be summarized in a phylogenetic tree.

In a hands-on example, the students were provided a representative coral gene and they searched for the corresponding gene in other species in the NCBI database. They found the gene to be present in a wide variety of species, which means that it was highly conserved during evolution.
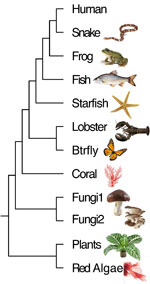
They also studied the phylogenetic tree that was generated from the gene data and learned several things that, at first glance may seem quite surprising, for example, the close relationship between lobsters and crabs (favorite food items in restaurants) and spiders and butterflies.
Surveys completed by the participants underlined that the event was well received and that the students look forward to more Science Saturdays at Rutgers University. As part of Rutgers Cooperative Extension, the 4-H Science program connects young people to Rutgers University faculty and their research. 4-H strives to inspire young people grades 5-12 to get involved in STEM.
January 2017


